
During the early stages of infection, the IP directs the synthesis of tas and bet mRNAs. HFV is also known to have a second promoter, termed the internal promoter (IP) located within the env gene, which drives tas and bet gene expression and is also transactivated by Tas protein. It encodes a protein that transactivates a long terminal repeat (LTR) promoter, which remains transcriptionally silent in the absence of Tas. The tas gene (transactivator of spumavirus) is however, known to be required for viral replication. Although a number of putative functions have been proposed for the bet gene, its role in vivo is still unknown. The HFV genome encodes the canonical retroviral gag, pol, and env genes, as well as at least two additional genes termed tas (bel-1) and bet. HFV particles shown budding from the ER (note immature particle appearance and characteristic envelope spikes)įoamy Viruses are considered complex retroviruses because they encode viral proteins that are not incorporated into viral particles. Like other Foamy Viruses, HFV has many characteristics that set it apart from the other well-known retroviruses, including a distinct morphology, budding from the endoplasmic reticulum rather than the plasma membrane and a unique replication strategy. 
There is also a lot of controversy about the criteria used to identify the isolate as HFV in most of these early studies, and so it may be that HFV is really an orphan virus and is just coincidentally associated with these disorders. Although it has been isolated from patients with various neoplastic and degenerative diseases such as myasthenia gravis, multiple sclerosis, thyroditis de Quervain and Graves disease, an etiological role for the virus has yet to be identified and little is known about its prevalence in human populations. HFV belongs to the genus of spumaviruses. troglodytes schweinfurthii in Kenya, the viruswas probably acquired as a zoonotic infection. Since the original HFV isolate came from a man who might have had contact with P. Recently, sequence comparisons between the original SFVcpz(hu) isolate and SFV from four distinct subspecies of chimpanzee demonstrate that SFVcpz(hu) is most closely related to foamy viruses found in Pan troglodytes schweinfurthii, whose natural habitat includes Kenya. The origin of SFVcpz(hu) was debated until 1994, when SFVcpz was cloned and sequenced the 86 to 95% amino acid identity between SFVcpz and SFVcpz(hu) suggested that SFVcpz(hu) is likely a variant of SFVcpz and not a unique isolate. At the time it was called SFVcpz(hu) because of its origin and its marked similarity to the Simian Foamy Virus found in chimpanzees. 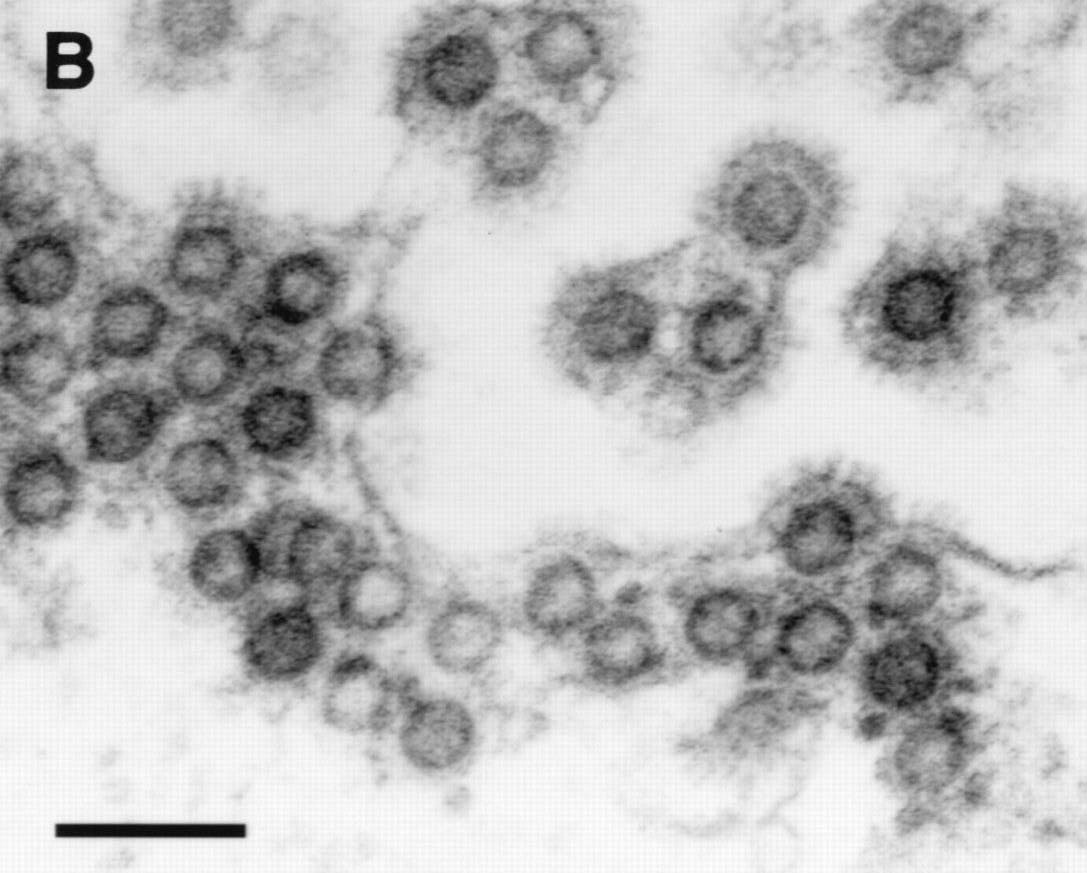
The human foamy virus (HFV) was firstisolated and identified in 1971 from cells released from a human nasopharyngealcarcinoma (NPC) in a Kenyan patient.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
